Towards Ultra-high-resolution E3SM Land Modeling on Exascale Computers
In December 2023, the manuscript titled “Towards Ultrahigh-resolution E3SM Land Modeling on Exascale Computers” was honored with the 2022 Best Paper Award from IEEE Computing in Science & Engineering, as recognized by the IEEE Computer Society Publications Board.
The refactorization of large-scale scientific code, such as the E3SM Land Model (ELM) for Exascale computers with hybrid computing architecture, poses a significant challenge for the scientific community. This paper introduces a holistic approach to the development of the ultrahigh-resolution ELM (uELM) over the Exascale computers at the DOE’s national laboratories. It also serves as a valuable reference for broad kilometer scale land modeling activities. The methodology commences with an analysis of the ELM software for computational model design, advances to the creation of porting tools for autonomous accelerator-ready code generation, and ends with preliminary uELM simulations over North America using an observation-constraint 1km x 1km forcing dataset[1].
This report summarizes the core concepts discussed in the award-winning paper and highlights the key milestones in the uELM development since the previous report in 2021.
Background
The U.S. Department of Energy (DOE) Energy Exascale Earth System Model (E3SM) project was conceived from the confluence of energy mission needs and disruptive changes in scientific computing technology. E3SM aims to optimize the use of DOE resources to meet the science needs of DOE with its unprecedented modeling capability at kilometer scales. The development of this kilometer-scale land model is a concurrent endeavor, in alignment with other E3SM components designed for Exascale computers, such as the kilometer-scale Atmosphere and Ocean models.
Ultrahigh-resolution E3SM Land Model (uELM) Development
ELM is a data-centric application with unique software characteristics
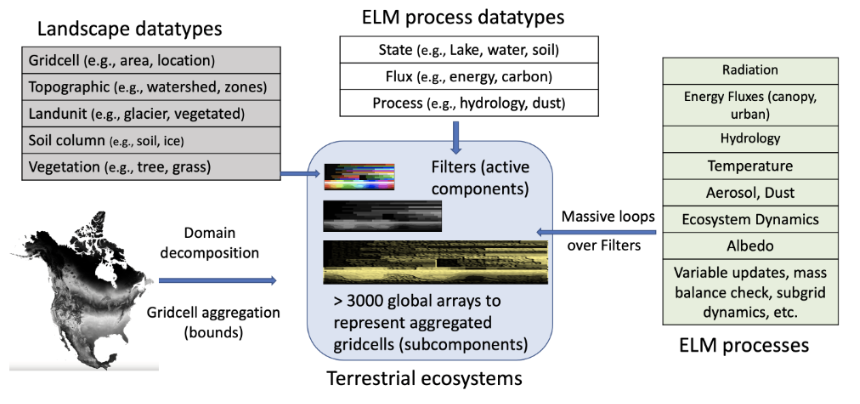
ELM emulates numerous biophysical and biochemical processes, representing the states and fluxes of energy, water, carbon, nitrogen, and phosphorus on individual land surface units, also known as gridcells. Each gridcell consists of subgrid elements, structured using a nested hierarchy of sub-grid components. Highly specialized landscape datatypes, including gridcell, topographic unit, land cover, soil column, and vegetation, are used to depict the heterogeneity of the Earth’s surface and subsurface. From a software standpoint, ELM is a data-centric, gridcell-independent terrestrial ecosystem simulation that involves extensive computational loops over gridcells and their subgrid components (Figure 1).
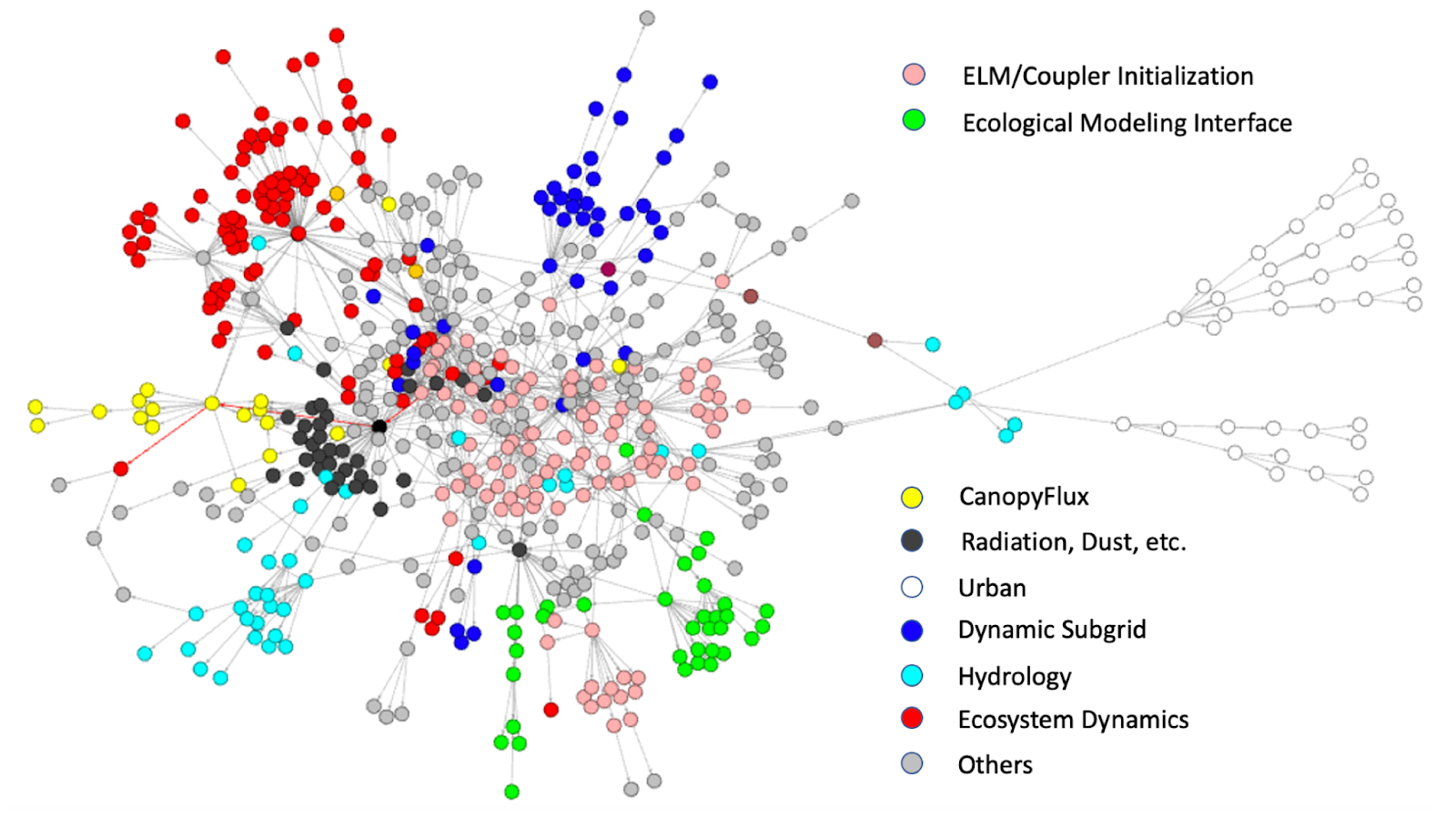
ELM possesses over 1000 subroutines, only a few of which are computationally intensive. Figure 2 depicts the primary functionality and the general functional call flow within each gridcell ELM simulation (data gathered with XScan [2]). The colors represent a selection of ELM submodels, including CanopyFlux, Hydrology, Urban, and Ecosystem Dynamics.
Computational Model for uELM Simulation on Hydrid Computing Architecture
A hybrid computational model for the uELM simulation was developed and integrated within the existing E3SM framework [1]. The ELM interfaces with the E3SM coupler have been preserved for the exchange of mass and energy, and a coupler-bypass has been designed to directly read atmospheric forcing from the disk. The majority of the original ELM configuration within the Common Infrastructure for Modeling the Earth (CIME) framework has been retained, using the CIME scripts for the uELM configuration, build, and batch job submission. The design strategy involved staging the ELM forcing data into the host memory, followed by data transfer to accelerators for massively parallel simulations.
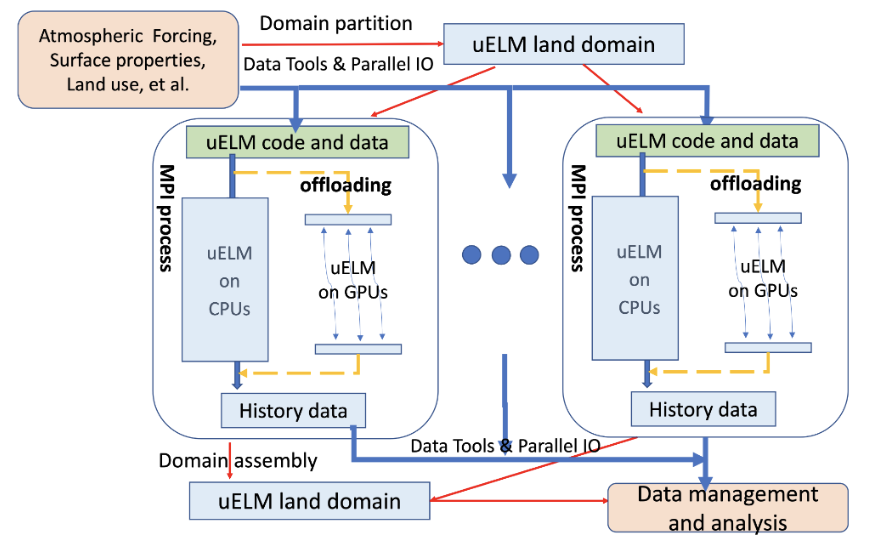
Currently, uELM is configured in the land-only mode within CIME (Figure 3). A data toolkit [3] has been developed to partition the uELM input dataset (e.g., forcing, surface properties, and land use) into subdomains using user-selected partition schemes (including the original round-robin partition scheme). uELM uses MPI for parallel execution, assigning a fixed number of grid cells to MPI processes through static domain decomposition. The metadata of domain partitions (indicated by red lines in Figure 3), such as the size and rank of subdomains, are collected to construct the entire uELM land domain, which is shared across the entire MPI communicator. On each MPI process, uELM uses parallel IO to read atmospheric forcing, uses the surface properties and land-use datasets (indicated by blue lines) to configure individual land cells, and subsequently conducts massively parallel simulations over these gridcells within each subdomain independently. Depending on the simulation domain’s size and the computer hardware, uELM can be executed on either CPUs or GPUs with OpenACC[4,5,6]. The uELM simulation results on individual MPI processes are stored in files and further handled by the uELM data management and analysis component.
Systematic Code Development and Essential Toolkit Creation
1. Software Toolkit for Porting ELM (SPEL) and End2End Simulation
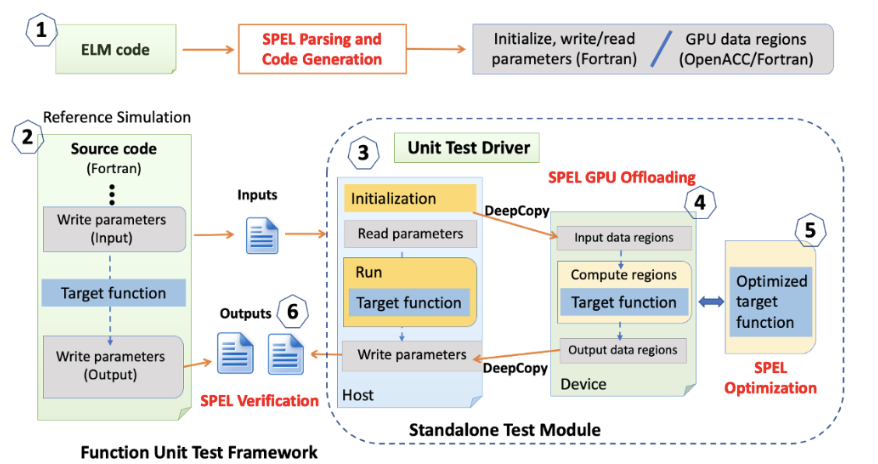
The scientists have designed a Python toolkit, named SPEL, to facilitate the porting of ELM modules onto Nvidia GPUs via OpenACC. SPEL is instrumental in generating 98% of the uELM code autonomously. The general process of code generation using SPEL is encapsulated in Figure 4. The SPEL workflow, embedded within a Functional Unit Testing (FUT) framework[7,8], comprises six steps: 1) SPEL parses the ELM code and generates a complete list of ELM function (global) parameters. For each individual ELM function, SPEL marks the active parameters and generates Fortran modules for reading, writing, initializing, and offloading parameters. 2) SPEL inserts the write modules before and after a specific ELM function to gather the input and output parameters from a reference ELM simulation. 3) SPEL constructs a unit test driver for standalone ELM module testing. This driver initializes and reads function parameters, executes the target ELM module, and saves the output. 4) SPEL generates GPU-ready ELM test modules with OpenACC directives. 5) SPEL optimizes the GPU-ready test module, including memory reduction, parallel loop, and data clauses. 6) SPEL verifies code accuracy at various stages of the ELM module testing, including CPU, GPU, and GPU-optimized stages.
2. End-to-end uELM code development and performance tuning
While the unit testing of uELM through SPEL lays a solid foundation for comprehensive code development on GPUs, the end-to-end uELM code development is still quite challenging. Four common issues were encountered during the process with the Nvidia HPC Software Development Kit: 1) The function that passes the array index to subroutines is flawed. It necessitates restructuring the routines using the index and explicitly passing the array elements as arguments. 2) The atomic operations (atomic directive) are inconsistent, despite potentially functioning within the FUT environment. It is crucial to ensure these atomic operations are performed over primitive datatypes. 3) The most frequently encountered error is a fatal one, which indicates that the variable, whether local or global, is only partially present on the device. Given the lack of direct access to the underlying algorithms and libraries, the scientists were forced to investigate various strategies to bypass this issue. And 4) The CUDA Debugger (within NVHPC) does not operate as efficiently as its CPU equivalents and requires excessive GPU memory. As a result, the scientists automated the creation of variable checking and verification functions (in FORTRAN) using Python scripts.
The simulation achieved a threefold speed increase on a fully loaded computing node of Summit. On a single Summit node, the time for 1 year of uELM runs over 36,000 gridcells (using CPU) is approximately 2.5 hours, with each timestep taking about 1 second. The execution time on GPUs for the same 1-year simulation is closer to 0.8 hours.
Portable and Collaborative Software Environment
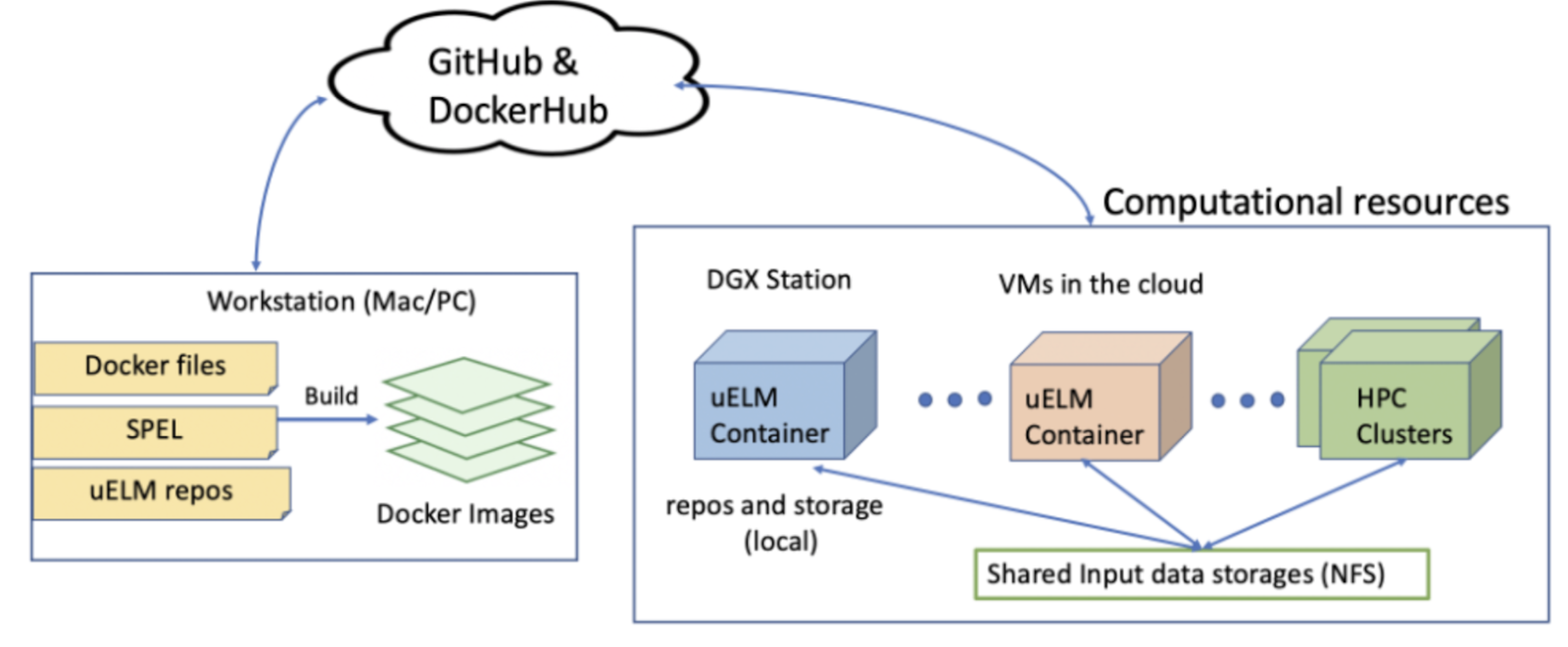
To facilitate community-based uELM developments with GPUs, the scientists have also created a portable, standalone software environment preconfigured with exemplar uELM input datasets, simulation cases, and source code. This environment (Figure 5), using Docker, encompasses all essential code, libraries, and system software for uELM development on GPUs. It also features a functional unit test framework and an offline model testbed for site-based ELM experiments[9].
High-Fidelity uELM Simulation over North America and Its Early Application
Computational Domain and High-Resolution Datasets
The computational domain (Figure 6, left panel) encompasses over 63 million (1km x 1km) gridcells, with a land mask comprising approximately 21.5 million gridcells. This quantity is roughly 350 times greater than the number of land gridcells (0.5-degree x 0.5-degree) displayed in the right panel.
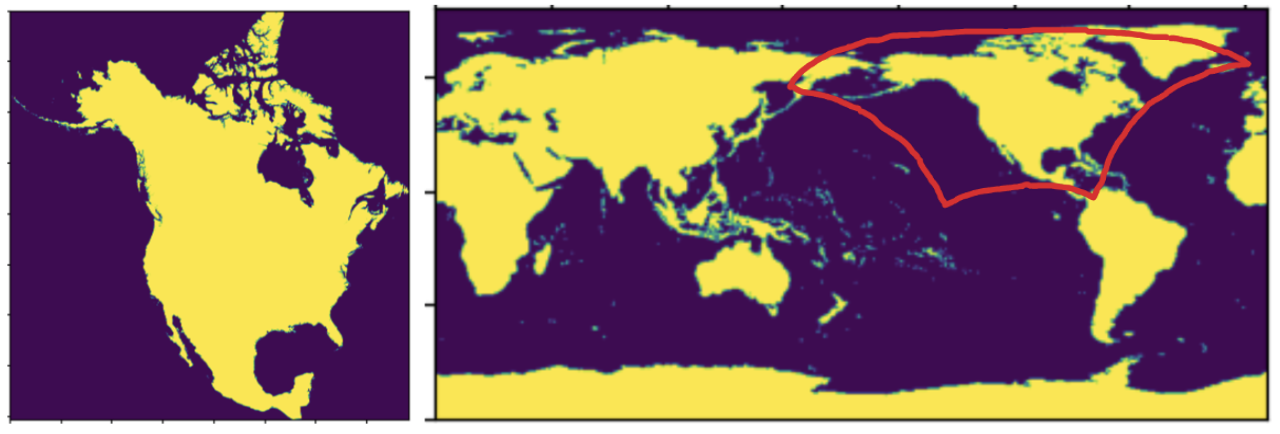
Figure 6: North America land mask (1km x 1km resolution) presented in a Lambert Conformal Conic (LCC) projection (left) and a global lank mask in WGS84 coordinates (0.5-degree x 0.5-degree) (right). The redline zone represents the rectangular domain in the left panel.
In this study, the scientists used temporally downscaled (3-hourly) meteorological forcing data from Daymet, which provides gridded estimates of daily weather parameters at 1 km spatial resolution from 1980 to the present for North America and Hawaii, and from 1950 to the present for Puerto Rico[10,11]. The Daymet dataset over North America, downscaled to 3-hourly, 1km by 1km resolution products, occupies roughly 40TB of disk space after 9 levels of data compression. Figure 7 shows samples of 1km by 1km atmospheric forcing at near-surface across North America.
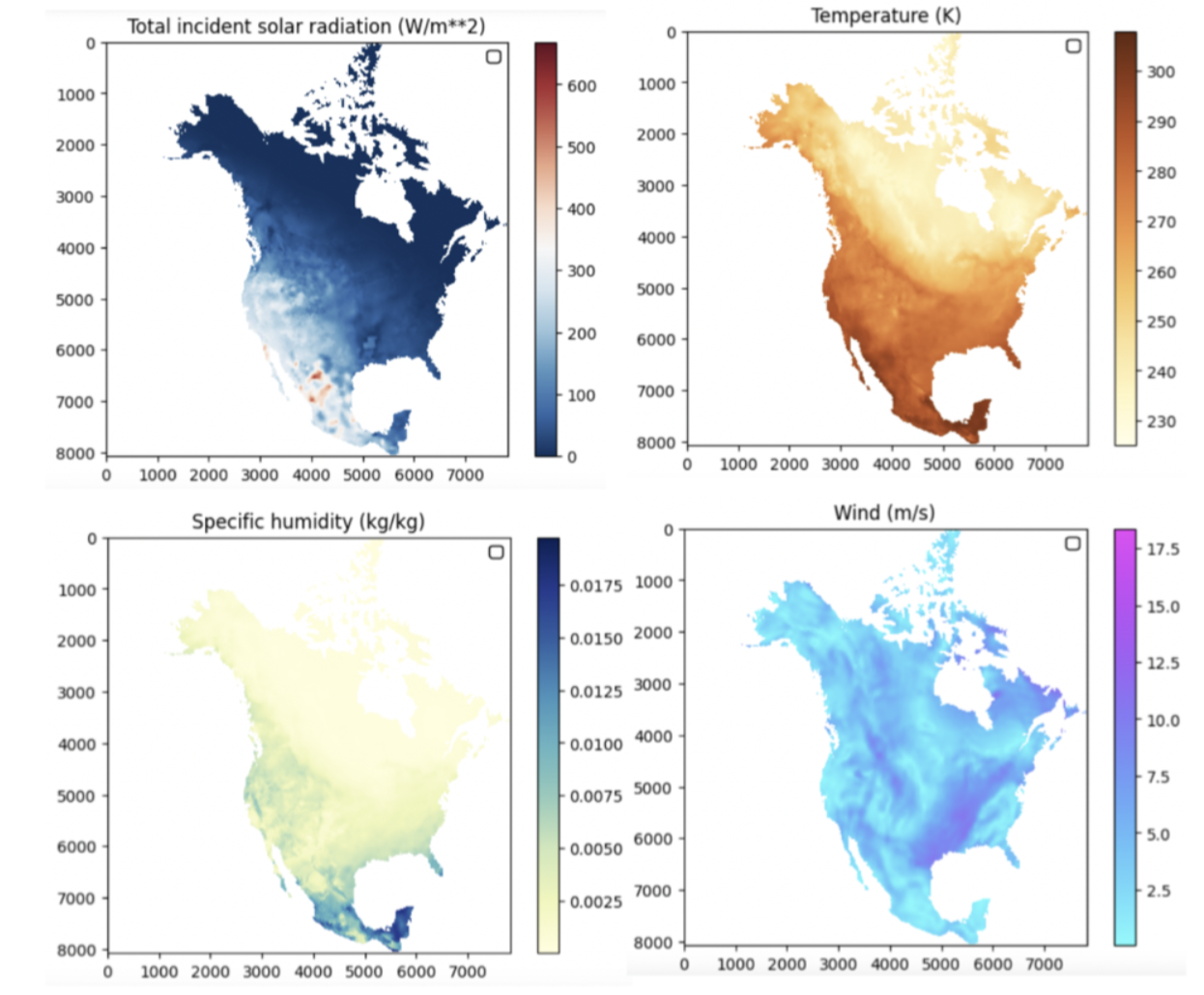
Figure 7. Samples of 3-hourly, high-resolution atmospheric forcing (radiation, temperature, specific humidity, and wind) across North America in January 2014.
For the purpose of code development, an efficient toolkit[3] was developed that creates several 1km-by-1km resolution ELM input datasets (over Daymet domain) from 0.5-degree by 0.5-degree global datasets (including surface properties, land use/land change, and nitrogen deposition). To further improve the quality of the dataset, the scientists are collaborating with other groups to integrate state-of-the-art data products (e.g., the fraction of naturally vegetated land and soil depth) and new landscape features (e.g., the maximal and minimal fraction of inundation and new features of urban area) into these 1km x 1km input dataset over North America.
An Early uELM Application over the Seward Peninsula, AK.
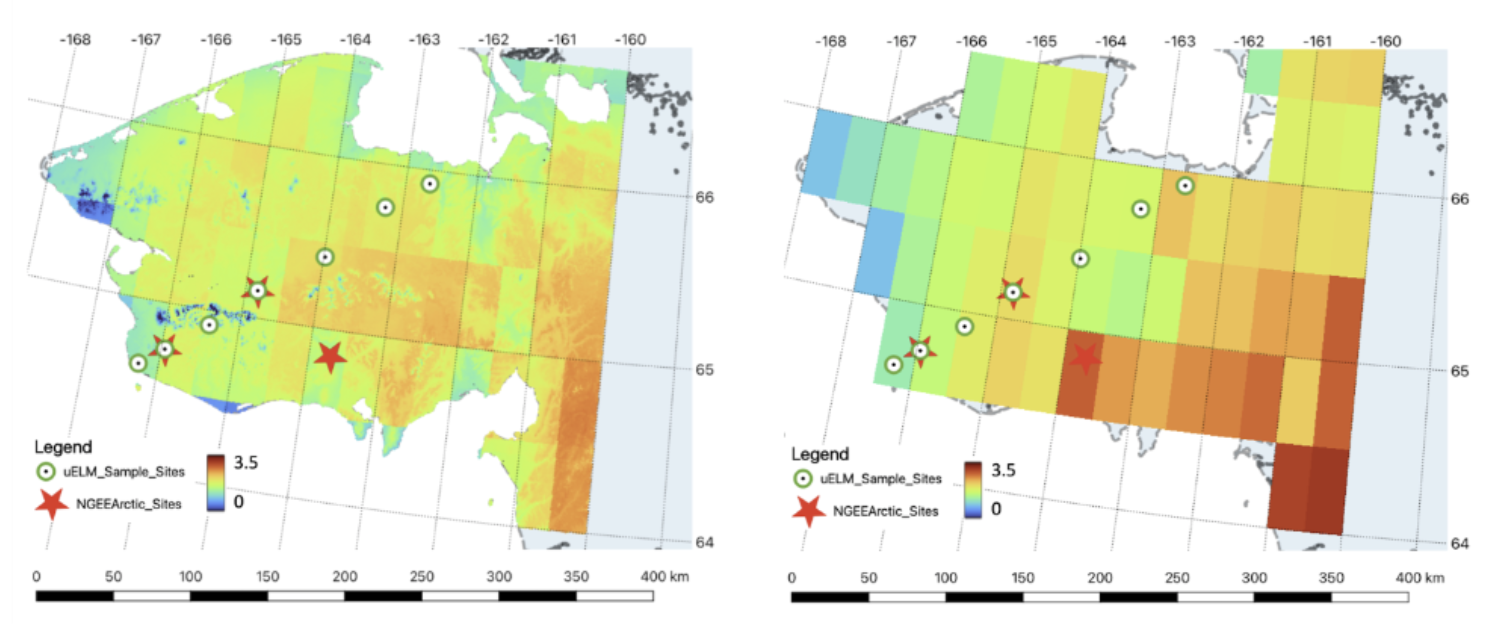
An uELM simulation was conducted over the Seward Peninsula at a resolution of 1km x 1km, which is the first experiment of its kind and yielded unprecedented results (Figure 8). The uELM simulation at the 1km-by-1km resolution enabled direct model validation and verification at similar scales as field experiments and site observation measurements[12].
Future Directions
Ultrahigh-resolution ELM simulations over North America require GPU-ready uELM simulation using many GPUs within world-class supercomputers. The IO operations and data exchange through E3SM coupler require further customization and tuning. Currently, scientists are evaluating several partition methods to split the massive input datasets into smaller subdomains suitable for parallel uELM simulations on compute nodes simultaneously. They are testing the scalability of GPU-ready ELM code on world-class supercomputers for large-scale uELM simulations. Additionally, scientists are investigating innovative strategies that employ neural network-based surrogate models to accelerate the uELM initialization.
References
- Wang, D., Schwartz, P., Yuan, F., Thornton, P., & Zheng, W. (2022). Toward Ultrahigh-Resolution E3SM Land Modeling on Exascale Computers. IEEE Computing in Science & Engineering, 24(6), 44-53. (Best Paper Award, 2022). DOI: 10.1109/MCSE.2022.3218990
- Zheng, W., Wang, D., & Song, F. (2019). XScan: an integrated tool for understanding open source community-based scientific code. In International Conference on Computational Science (pp. 226-237). Lecture Notes in Computer Science, Springer, Cham. DOI: 10.1007/978-3-030-22734-0_17
- Wang, D., Yuan, F., Schwartz, P., Kao, S., Thornton, M., Ricciuto, D., Thornton, P., & Reis, S. (2023). Data Toolkit for Ultrahigh-resolution ELM Data Partition and Generation. In Proceedings of 25th Congress on Modeling and Simulation, pp 996-1003, https://modsim2023.exordo.com/files/papers/46/final_draft/wang46.pdf
- Schwartz, P., Wang, D., Yuan, F., & Thornton, P. (2022). SPEL: Software Tool for Porting E3SM Land Model with OpenACC in a Functional Unit Test Framework. In the 9th Workshop on Accelerator Programming Using Directives. DOI: 10.1109/WACCPD56842.2022.00010.
- Schwartz, P., Wang, D., Yuan, F., & Thornton, P. (2023). Developing Ultrahigh-Resolution E3SM Land Model for GPU Systems. In Gervasi et al. (Eds.), Computational Science and Its Applications – ICCSA 2023 (pp. 13956). Lecture Notes in Computer Science. Springer, Cham. DOI: 10.1007/978-3-031-36805-9_19
- Schwartz, P., Wang, D., Yuan, F., & Thornton, P. (2022). Developing an ELM Ecosystem Dynamics Model on GPU with OpenACC. In Lecture Notes on Computer Science, D. Groen et al. (Eds.): ICCS 2022, LNCS 13351, 1–13. DOI:10.1007/978-3-031-08754-7_38
- Wang, D., Wu, W., Janjusic, T., Xu, Y., Iversen, C., Thornton, P., & Krassovisk, M. (2015, May). Scientific functional testing platform for environmental models: An application to community land model. In International Workshop on Software Engineering for High Performance Computing in Science, 37th International Conference on Software Engineering. DOI:10.1109/SE4HPCS.2015.10
- Wang, D., Xu, Y., Thornton, P., King, A., Gu, L., Steed, C., & Schuchart. J. (2014). A functional testing platform for the community land model. Environmental Modeling and Software, 55:25-31. DOI: 10.1016/j.envsoft.2014.01.015
- Wang, D., Schwartz, P., Yuan, F., Ricciuto, D., Thornton, P., Layton, C., Eaglebarge, F., & Cao, Q. (2024). Portable Software Environment for Ultrahigh Resolution ELM Modeling on GPUs. (in review)
- Thornton, P. E., Shrestha, R., Thornton, M., Kao, S.-C., Wei, Y., & Wilson, B. E. (2021). Gridded daily weather data for North America with comprehensive uncertainty quantification. Scientific Data, 8(1), 190. https://doi.org/10.1038/s41597-021-00973-0
- Thornton, M. M., Shrestha, R., Wei, Y., Thornton, P. E., Kao, S.-C., & Wilson, B. E. (2020). Daymet: Daily Surface Weather Data on a 1-km Grid for North America, Version 4. ORNL DAAC. Daymet: Daily Surface Weather Data on a 1-km Grid for North America, Version 4, https://doi.org/10.3334/ORNLDAAC/1840
- Yuan, F., Wang, D., Kao, S., Thornton, M., Ricciuto, D., Sulmon, V., Iversen, C., Schwartz, P., & Thornton, P. (2023). An ultrahigh-resolution E3SM Land Model simulation framework and its first application to the Seward Peninsula in Alaska, Journal of Computational Science, Vol73, pp02145. DOI: 10.1016/j.jocs.2023.102145
Funding
- The U.S. Department of Energy Office of Science, Biological and Environmental Research supported this research as part of the Earth System Model Development Program Area through the Energy Exascale Earth System Model (E3SM) project. This research used resources of the Oak Ridge Leadership Computing Facility and Computing and Data Environment for Science (CADES) at the Oak Ridge National Laboratory, which are supported by the DOE’s Office of Science under Contract No. DE-AC05-00OR22725.
Contacts
- Dali Wang, Oak Ridge National Laboratory
- Peter Schwartz, Oak Ridge National Laboratory
- Peter Thornton, Oak Ridge National Laboratory
This article is a part of the E3SM “Floating Points” Newsletter, to read the full Newsletter check: